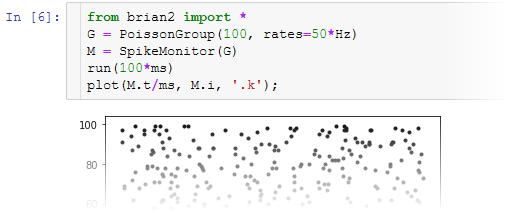
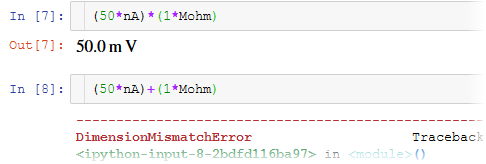
Why use Brian?
Download and installation
conda install -c conda-forge brian2or
pip install brian2
For more details, see the installation instructions in the documentation.
Source code available on GitHub.
News
2026-03-16
GSoC 2026
2025-12-10
New release: Brian 2.10
2025-05-16
New release: Brian 2.9.0
2025-03-10
GSoC 2025
Articles
2021-09-24
Bug hunt episode 2: a strange file appears
2020-10-08
Getting the timing right (scheduling 2)
2020-09-04
Getting the timing right (scheduling 1)
Community
Community forum
Chat group
Papers using Brian
Software ecosystem
Models (and older ones)
How to contribute
Team
-
Romain Brette
ISIR, Sorbonne University, INSERM, Paris -
Ben Evans
University of Sussex -
Dan Goodman
Imperial College London -
Marcel Stimberg
ISIR, Sorbonne University, Paris
For a full list of contributors, see the AUTHORS file in the repository. All contributions (not only in the form of code) are listed in the release notes.